
- #.BAM FILE FORMAT ARCHIVE#
- #.BAM FILE FORMAT FULL#
- #.BAM FILE FORMAT SOFTWARE#
- #.BAM FILE FORMAT PLUS#
- #.BAM FILE FORMAT SERIES#
We also make the collation algorithm available in the form of an API for other projects. Using this algorithm tasks like duplicate marking in BAM files and conversion of BAM files to the FastQ format can be performed very efficiently with limited resources. The employed collation algorithm avoids time and space consuming sorting of alignments by read name where this is possible without using more than a specified amount of main memory. In this paper we introduce biobambam, a set of tools based on the efficient collation of alignments in BAM files by read name.
#.BAM FILE FORMAT FULL#
In this order paired reads are usually separated in the file, which complicates some other applications like duplicate marking or conversion to the FastQ format which require to access the full information of the pairs.
#.BAM FILE FORMAT PLUS#
See the spec for a detailed list of commonly used tags and what they mean.Sequence alignment data is often ordered by coordinate (id of the reference sequence plus position on the sequence where the fragment was mapped) when stored in BAM files, as this simplifies the extraction of variants between the mapped data and the reference or of variants within the mapped data. A bunch of different information can be stored here and they appear as key/value pairs. Additional optional information is also contained within the alignment, TAGs.(col 11) the query quality for this alignment, QUAL, one for each base in the query sequence.(col 10) the query sequence for this alignment, SEQ.(col 9) length of this group from the leftmost position to the rightmost position, ISIZE or TLEN.Beware to always use the correct base when referencing positions. For SAM, the reference starts at 1, so this value is 1-based, while for BAM the reference starts at 0,so this value is 0-based. (col 8) leftmost position of where the next alignment in this group maps to the reference, MPOS or PNEXT.(A group is alignments with the same query name.) In paired alignments, it is the mate's reference sequence name. (col 7) the reference sequence name of the next alignment in this group, MRNM or RNEXT.(col 6) string indicating alignment information that allows the storing of clipped, CIGAR.(col 5) mapping quality, MAPQ, which contains the "phred-scaled posterior probability that the mapping position" is wrong.(col 4) leftmost position of where this alignment maps to the reference, POS.(col 3) reference sequence name(ref), RNAME, often contains the Chromosome name( chr#).is this read a PCR or optical duplicate?.is the next fragment the reverse strand?.a bitwise set of information describing the alignment, FLAG.It is used to group/identify alignments that are together, like paired alignments or a read that appears in multiple alignments. query name, QNAME (SAM)/read_name (BAM).What Information Does SAM/BAM Have for an Alignment The alignment section contains the information for each sequence about where/how it aligns to the reference genome. The alignments then associate themselves with specific header information. The header section may contain information about the entire file and additional information for alignments. Refer to the specs to see a format description.īoth SAM & BAM files contain an optional header section followed by the alignment section.
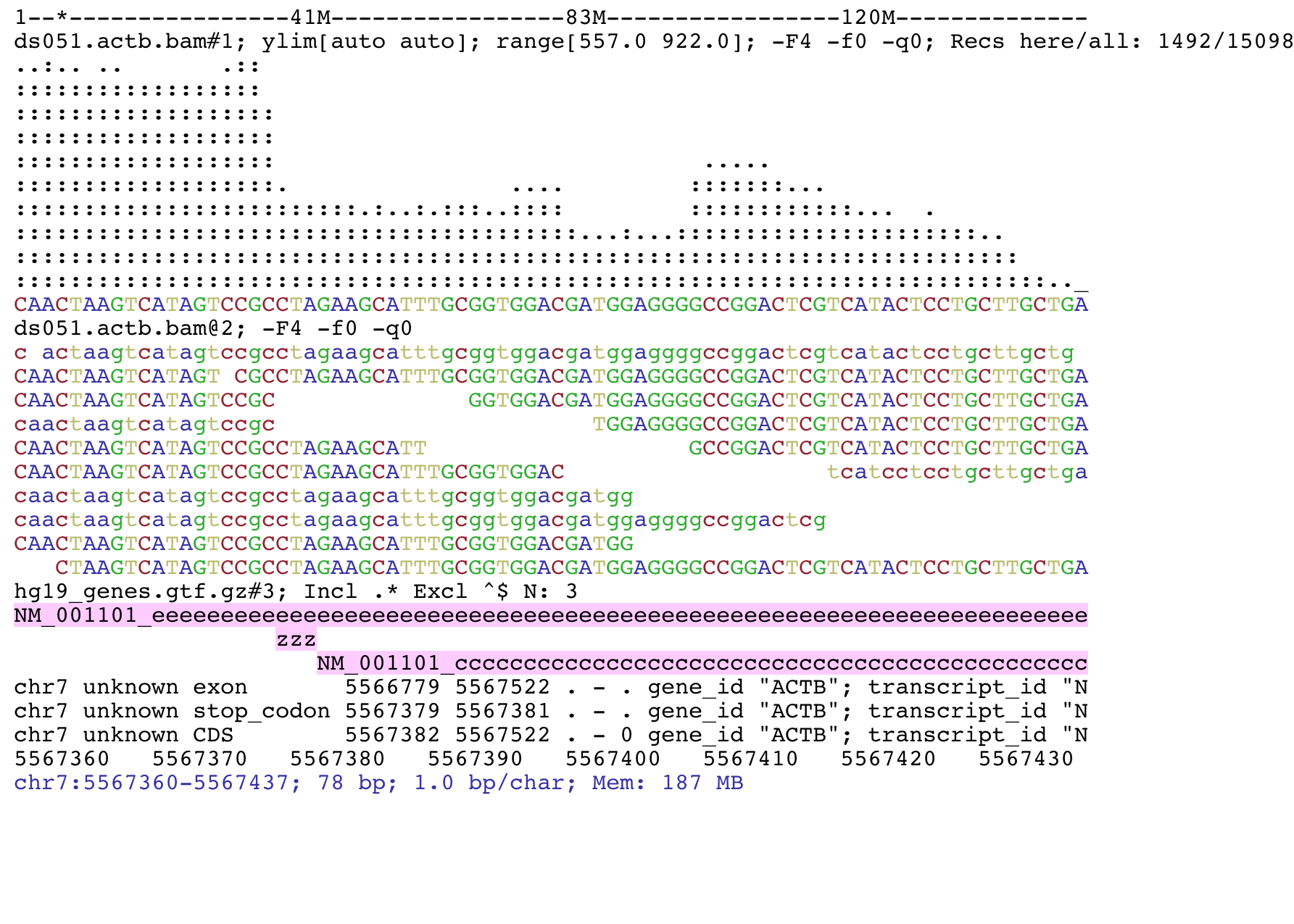
SAM files and BAM files contain the same information, but in a different format.
#.BAM FILE FORMAT SOFTWARE#
If you are writing software to read SAM or BAM data, our C++ libStatGen is a good resource to use. The current definition of the format is at.
#.BAM FILE FORMAT ARCHIVE#
In the future, SAM will also be used to archive unaligned sequence data generated directly from sequencing machines. Most often it is generated as a human readable version of its sister BAM format, which stores the same data in a compressed, indexed, binary form.Ĭurrently, most SAM format data is output from aligners that read FASTQ files and assign the sequences to a position with respect to a known reference genome.
#.BAM FILE FORMAT SERIES#
The SAM Format is a text format for storing sequence data in a series of tab delimited ASCII columns.


 0 kommentar(er)
0 kommentar(er)
